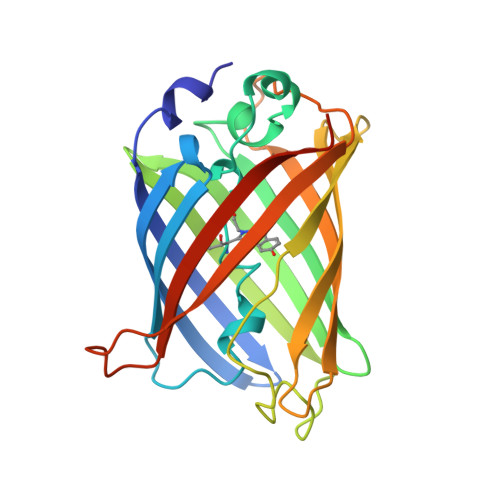
Structural basis for dual excitation and photoisomerization of the Aequorea victoria green fluorescent protein.
Brejc, K., Sixma, T.K., Kitts, P.A., Kain, S.R., Tsien, R.Y., Ormo, M., Remington, S.J.(1997) Proc Natl Acad Sci U S A 94: 2306-2311
- PubMed: 9122190
- DOI: https://doi.org/10.1073/pnas.94.6.2306
- Primary Citation of Related Structures:
1EMB - PubMed Abstract:
The 2.1-A resolution crystal structure of wild-type green fluorescent protein and comparison of it with the recently determined structure of the Ser-65 --> Thr (S65T) mutant explains the dual wavelength absorption and photoisomerization properties of the wild-type protein. The two absorption maxima are caused by a change in the ionization state of the chromophore. The equilibrium between these states appears to be governed by a hydrogen bond network that permits proton transfer between the chromophore and neighboring side chains. The predominant neutral form of the fluorophore maximally absorbs at 395 nm. It is maintained by the carboxylate of Glu-222 through electrostatic repulsion and hydrogen bonding via a bound water molecule and Ser-205. The ionized form of the fluorophore, absorbing at 475 nm, is present in a minor fraction of the native protein. Glu-222 donates its charge to the fluorophore by proton abstraction through a hydrogen bond network, involving Ser-205 and bound water. Further stabilization of the ionized state of the fluorophore occurs through a rearrangement of the side chains of Thr-203 and His-148. UV irradiation shifts the ratio of the two absorption maxima by pumping a proton relay from the neutral chromophore's excited state to Glu-222. Loss of the Ser-205-Glu-222 hydrogen bond and isomerization of neutral Glu-222 explains the slow return to the equilibrium dark-adapted state of the chromophore. In the S65T structure, steric hindrance by the extra methyl group stabilizes a hydrogen bonding network, which prevents ionization of Glu-222. Therefore the fluorophore is permanently ionized, causing only a 489-nm excitation peak. This new understanding of proton redistribution in green fluorescent protein should enable engineering of environmentally sensitive fluorescent indicators and UV-triggered fluorescent markers of protein diffusion and trafficking in living cells.
Organizational Affiliation:
Netherlands Cancer Institute, Department of Molecular Carcinogenesis, Amsterdam.