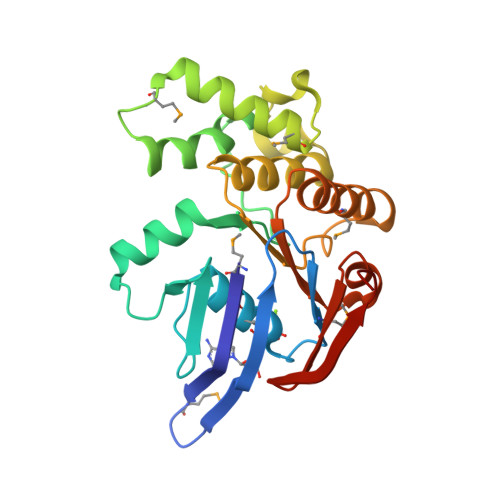
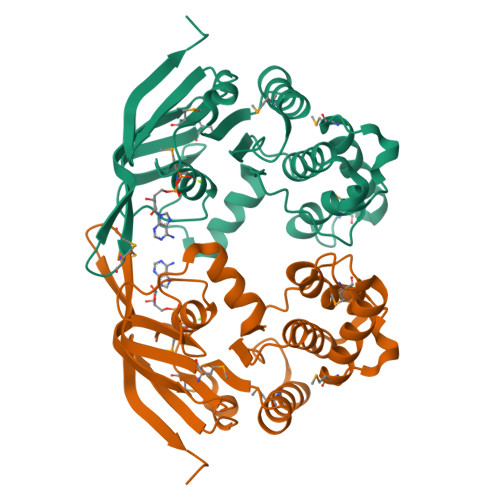
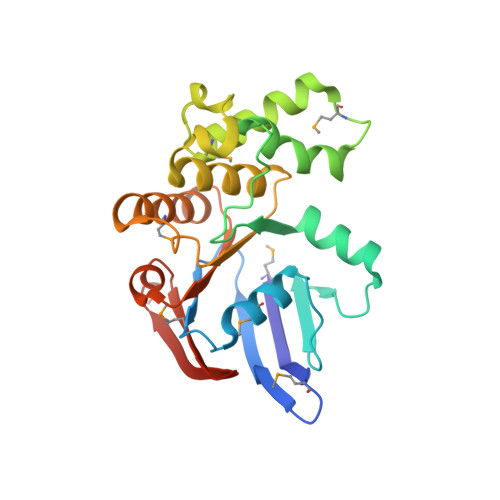
The crystal structure of the MJ0796 ATP-binding cassette. Implications for the structural consequences of ATP hydrolysis in the active site of an ABC transporter.
Yuan, Y.R., Blecker, S., Martsinkevich, O., Millen, L., Thomas, P.J., Hunt, J.F.(2001) J Biological Chem 276: 32313-32321
- PubMed: 11402022
- DOI: https://doi.org/10.1074/jbc.M100758200
- Primary Citation of Related Structures:
1F3O - PubMed Abstract:
The crystal structure of the MJ0796 ATP-binding cassette, a member of the o228/LolD transporter family, has been determined at 2.7-A resolution with MgADP bound at its active site. Comparing this structure with that of the ATP-bound form of the HisP ATP-binding cassette (Hung, L. W., Wang, I. X., Nikaido, K., Liu, P. Q., Ames, G. F., and Kim, S. H. (1998) Nature 396, 703-707) shows a 5-A withdrawal of a phylogenetically invariant glutamine residue from contact with the gamma-phosphate of ATP in the active site. This glutamine is located in a protein segment that links the rigid F(1)-type ATP-binding core of the enzyme to an ABC transporter-specific alpha-helical subdomain that moves substantially away from the active site in the MgADP-bound structure of MJ0796 compared with the ATP-bound structure of HisP. A similar conformational effect is observed in the MgADP-bound structure of MJ1267 (Karpowich, N., et al. (2001) Structure, in press), establishing the withdrawal of the glutamine and the coupled outward rotation of the alpha-helical subdomain as consistent consequences of gamma-phosphate release from the active site of the transporter. Considering this subdomain movement in the context of a leading model for the physiological dimer of cassettes present in ABC transporters indicates that it produces a modest mechanical change that is likely to play a role in facilitating nucleotide exchange out of the ATPase active site. Finally, it is noteworthy that one of the intersubunit packing interactions in the MJ0796 crystal involves antiparallel beta-type hydrogen bonding interactions between the outermost beta-strands in the two core beta-sheets, leading to their fusion into a single extended beta-sheet, a type of structural interaction that has been proposed to play a role in mediating the aggregation of beta-sheet-containing proteins.
Organizational Affiliation:
Department of Biological Sciences, Columbia University, New York, New York 10027, USA.