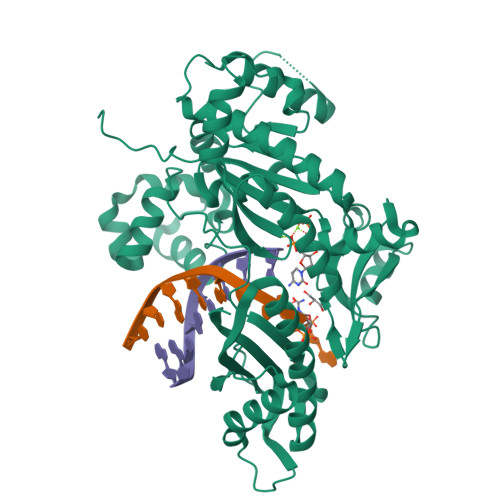
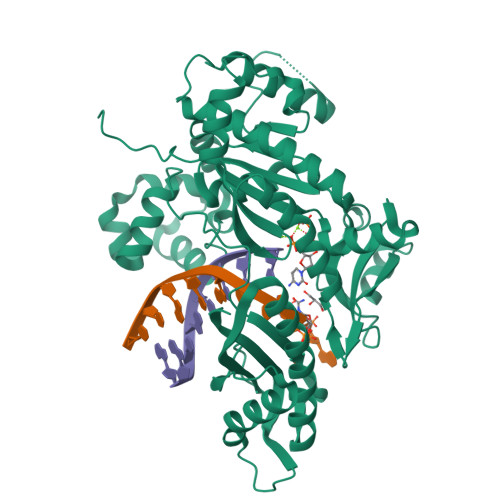
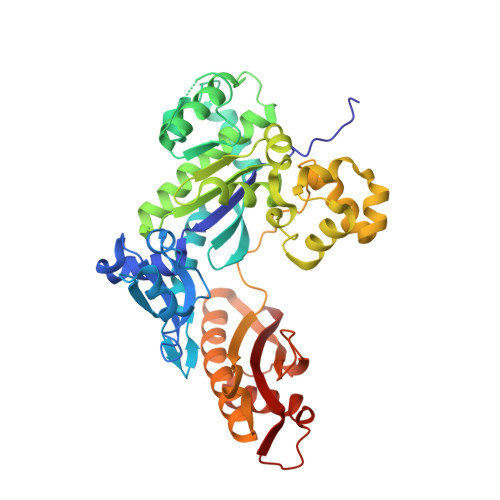
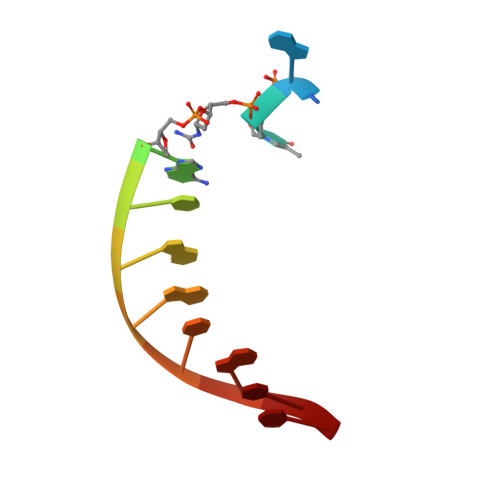
Replication Bypass of the N -(2-Deoxy-d-erythro-pentofuranosyl)-urea DNA Lesion by Human DNA Polymerase eta.
Tomar, R., Li, S., Egli, M., Stone, M.P.(2024) Biochemistry 63: 754-766
- PubMed: 38413007
- DOI: https://doi.org/10.1021/acs.biochem.3c00569
- Primary Citation of Related Structures:
8UJT, 8UJV, 8UJX, 8UK4 - PubMed Abstract:
Urea lesions in DNA arise from thymine glycol (Tg) or 8-oxo-dG; their genotoxicity is thought to arise in part due to their potential to accommodate the insertion of all four dNTPs during error-prone replication. Replication bypass with human DNA polymerase η (hPol η) confirmed that all four dNTPs were inserted opposite urea lesions but with purines exhibiting greater incorporation efficiency. X-ray crystal structures of ternary replication bypass complexes in the presence of Mg 2+ ions with incoming dNTP analogs dAMPnPP, dCMPnPP, dGMPnPP, and dTMPnPP bound opposite urea lesions (hPol η·DNA·dNMPnPP complexes) revealed all were accommodated by hPol η. In each, the Watson-Crick face of the dNMPnPP was paired with the urea lesion, exploiting the ability of the amine and carbonyl groups of the urea to act as H-bond donors or acceptors, respectively. With incoming dAMPnPP or dGMPnPP, the distance between the imino nitrogen of urea and the N9 atoms of incoming dNMPnPP approximated the canonical distance of 9 Å in B-DNA. With incoming dCMPnPP or dTMPnPP, the corresponding distance of about 7 Å was less ideal. Improved base-stacking interactions were also observed with incoming purines vs pyrimidines. Nevertheless, in each instance, the α-phosphate of incoming dNMPnPPs was close to the 3'-hydroxyl group of the primer terminus, consistent with the catalysis of nucleotidyl transfer and the observation that all four nucleotides could be inserted opposite urea lesions. Preferential insertion of purines by hPol η may explain, in part, why the urea-directed spectrum of mutations arising from Tg vs 8-oxo-dG lesions differs.
Organizational Affiliation:
Department of Chemistry, Vanderbilt Ingram Cancer Center, and Vanderbilt Center for Structural Biology, Vanderbilt University, Nashville, Tennessee 37235, United States.