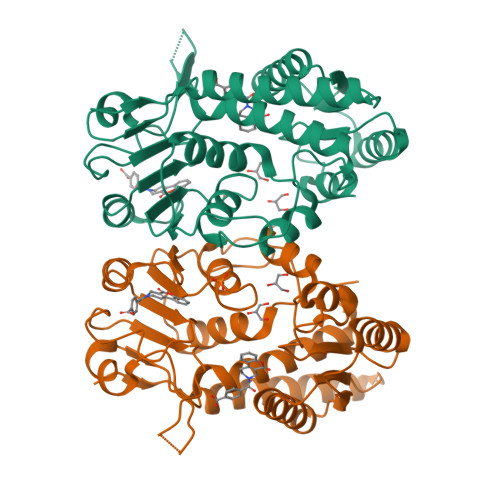
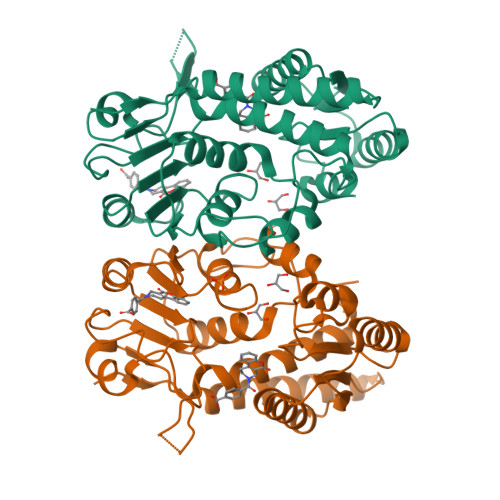
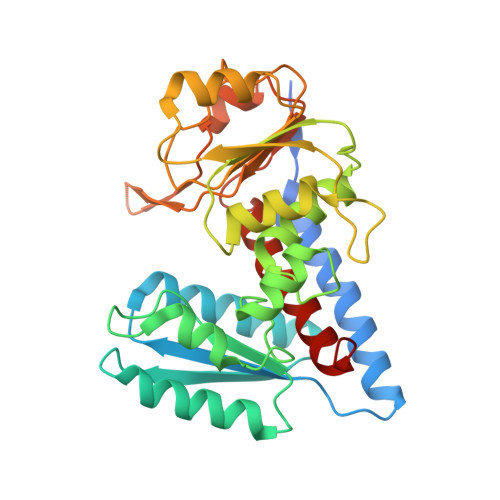
Structure-Based Design and Synthesis of an Isozyme-Selective MTHFD2 Inhibitor with a Tricyclic Coumarin Scaffold.
Kawai, J., Ota, M., Ohki, H., Toki, T., Suzuki, M., Shimada, T., Matsui, S., Inoue, H., Sugihara, C., Matsuhashi, N., Matsui, Y., Takaishi, S., Nakayama, K.(2019) ACS Med Chem Lett 10: 893-898
- PubMed: 31223444
- DOI: https://doi.org/10.1021/acsmedchemlett.9b00069
- Primary Citation of Related Structures:
6JIB, 6JID - PubMed Abstract:
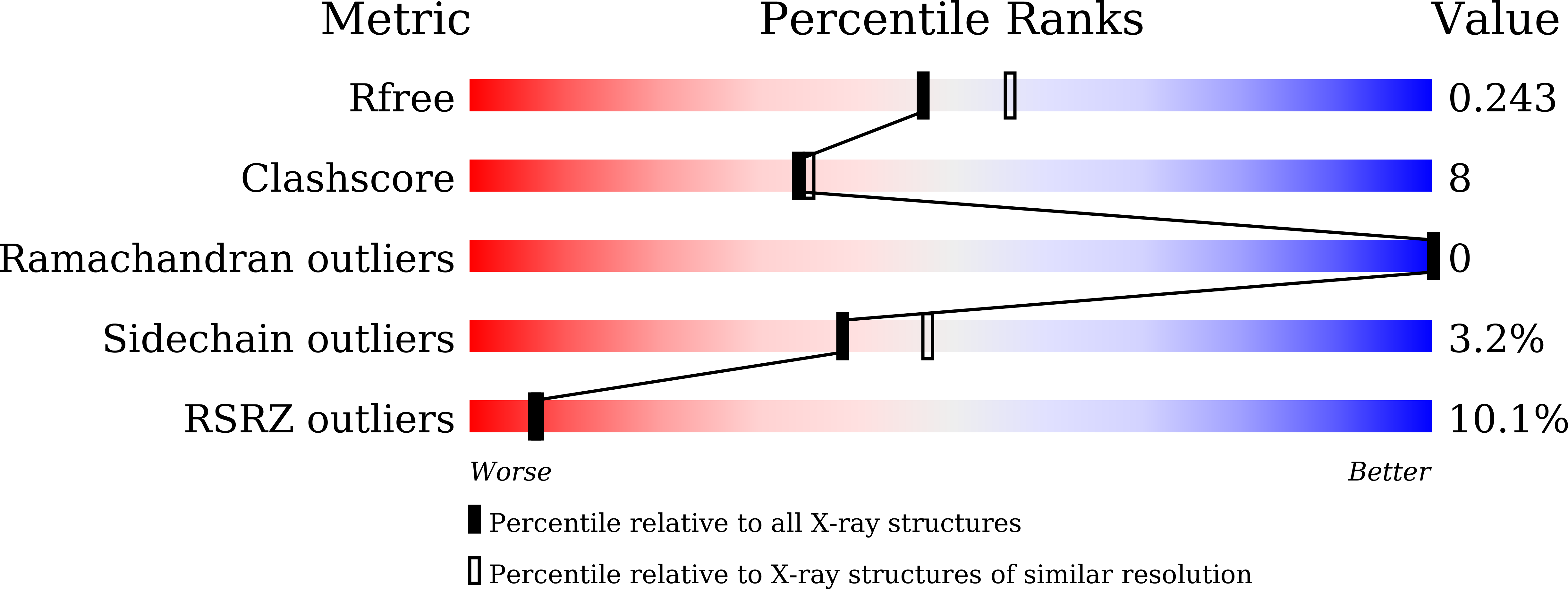
Methylenetetrahydrofolate dehydrogenase 2 (MTHFD2) plays a key role in one-carbon (1C) metabolism in human mitochondria, and its high expression correlates with poor survival of patients with various types of cancer. An isozyme-selective MTHFD2 inhibitor is highly attractive for potential use in cancer treatment. Herein, we disclose a novel isozyme-selective MTHFD2 inhibitor DS44960156, with a tricyclic coumarin scaffold, which was initially discovered via high-throughput screening (HTS) and improved using structure-based drug design (SBDD). DS44960156 would offer a good starting point for further optimization based on the following features: (1) unprecedented selectivity (>18-fold) for MTHFD2 over MTHFD1, (2) a molecular weight of less than 400, and (3) good ligand efficiency (LE).
Organizational Affiliation:
R&D Division, Daiichi Sankyo Co., Ltd., 1-2-58 Hiromachi, Shinagawa-ku, Tokyo 140-8710, Japan.