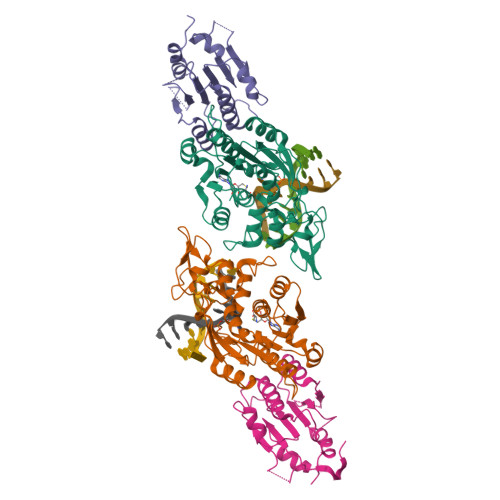
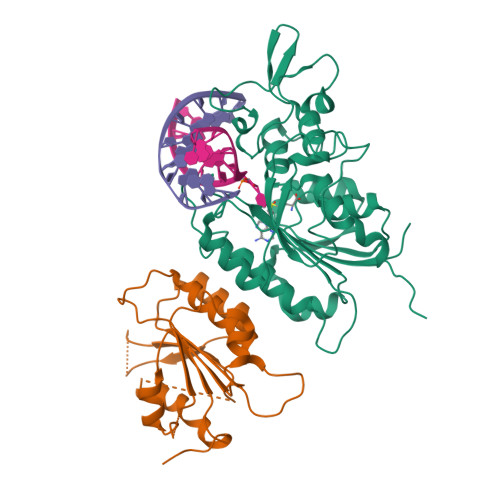
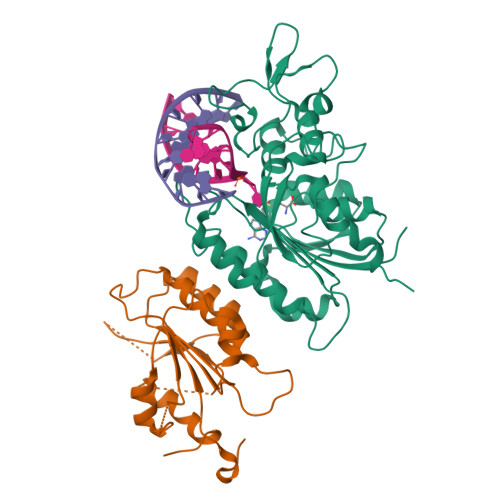
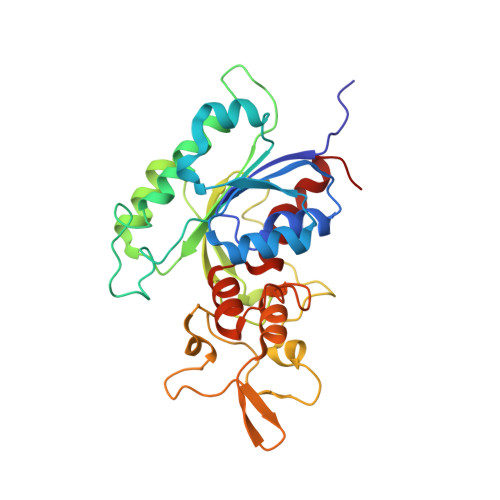
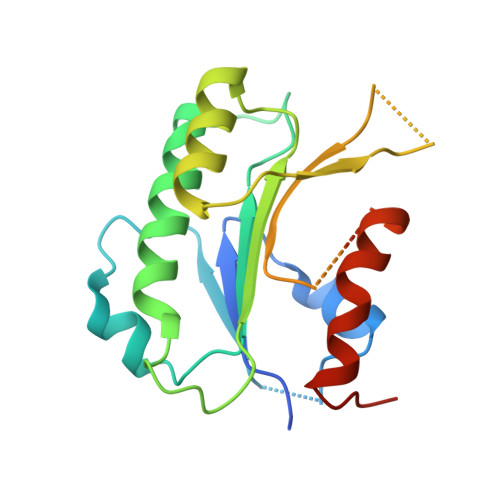
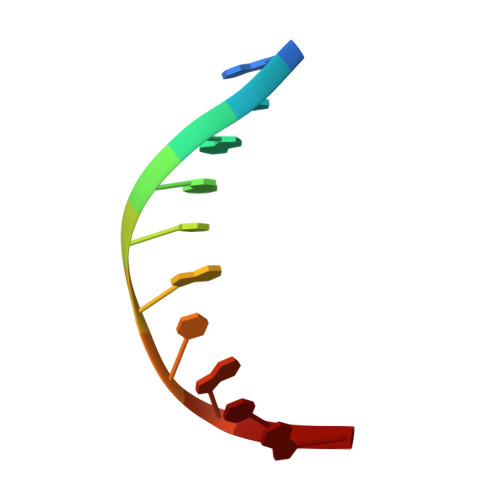
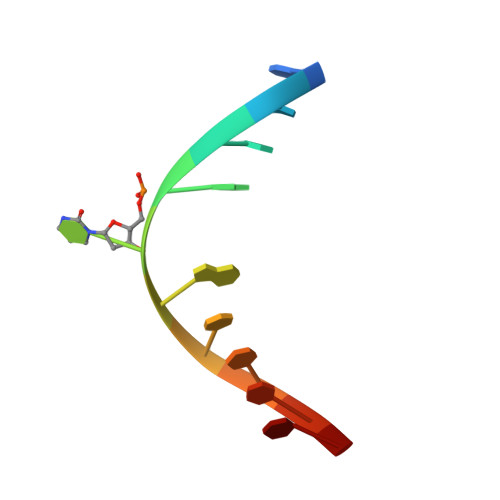
Structural basis for DNMT3A-mediated de novo DNA methylation.
Zhang, Z.M., Lu, R., Wang, P., Yu, Y., Chen, D., Gao, L., Liu, S., Ji, D., Rothbart, S.B., Wang, Y., Wang, G.G., Song, J.(2018) Nature 554: 387-391
- PubMed: 29414941
- DOI: https://doi.org/10.1038/nature25477
- Primary Citation of Related Structures:
5YX2, 6BRR, 6F57 - PubMed Abstract:
DNA methylation by de novo DNA methyltransferases 3A (DNMT3A) and 3B (DNMT3B) at cytosines is essential for genome regulation and development. Dysregulation of this process is implicated in various diseases, notably cancer. However, the mechanisms underlying DNMT3 substrate recognition and enzymatic specificity remain elusive. Here we report a 2.65-ångström crystal structure of the DNMT3A-DNMT3L-DNA complex in which two DNMT3A monomers simultaneously attack two cytosine-phosphate-guanine (CpG) dinucleotides, with the target sites separated by 14 base pairs within the same DNA duplex. The DNMT3A-DNA interaction involves a target recognition domain, a catalytic loop, and DNMT3A homodimeric interface. Arg836 of the target recognition domain makes crucial contacts with CpG, ensuring DNMT3A enzymatic preference towards CpG sites in cells. Haematological cancer-associated somatic mutations of the substrate-binding residues decrease DNMT3A activity, induce CpG hypomethylation, and promote transformation of haematopoietic cells. Together, our study reveals the mechanistic basis for DNMT3A-mediated DNA methylation and establishes its aetiological link to human disease.
Organizational Affiliation:
Department of Biochemistry, University of California, Riverside, California 92521, USA.