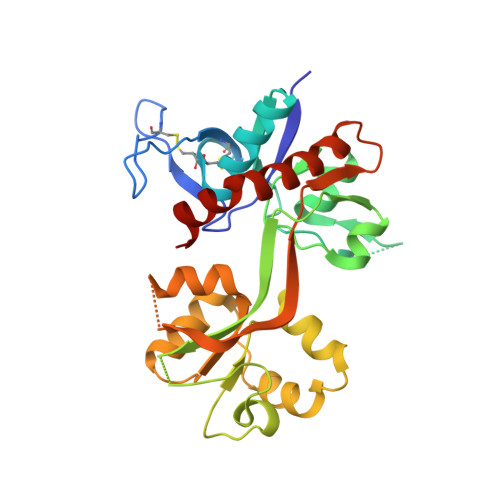
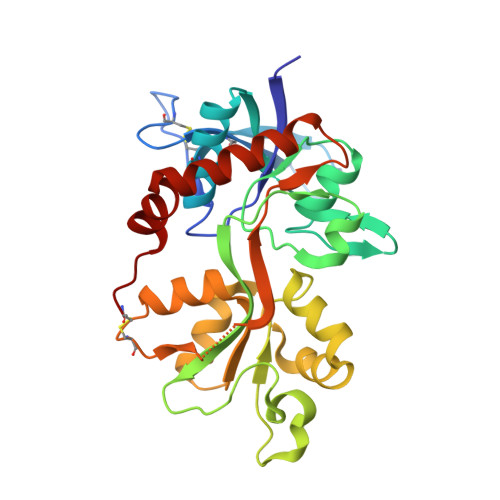
Structural Basis for Negative Allosteric Modulation of GluN2A-Containing NMDA Receptors.
Yi, F., Mou, T.C., Dorsett, K.N., Volkmann, R.A., Menniti, F.S., Sprang, S.R., Hansen, K.B.(2016) Neuron 91: 1316-1329
- PubMed: 27618671
- DOI: https://doi.org/10.1016/j.neuron.2016.08.014
- Primary Citation of Related Structures:
5I56, 5I57, 5I58, 5I59, 5JTY - PubMed Abstract:
NMDA receptors mediate excitatory synaptic transmission and regulate synaptic plasticity in the central nervous system, but their dysregulation is also implicated in numerous brain disorders. Here, we describe GluN2A-selective negative allosteric modulators (NAMs) that inhibit NMDA receptors by stabilizing the apo state of the GluN1 ligand-binding domain (LBD), which is incapable of triggering channel gating. We describe structural determinants of NAM binding in crystal structures of the GluN1/2A LBD heterodimer, and analyses of NAM-bound LBD structures corresponding to active and inhibited receptor states reveal a molecular switch in the modulatory binding site that mediate the allosteric inhibition. NAM binding causes displacement of a valine in GluN2A and the resulting steric effects can be mitigated by the transition from glycine bound to apo state of the GluN1 LBD. This work provides mechanistic insight to allosteric NMDA receptor inhibition, thereby facilitating the development of novel classes NMDA receptor modulators as therapeutic agents.
Organizational Affiliation:
Department of Biomedical and Pharmaceutical Sciences, University of Montana, Missoula, MT 59812, USA.