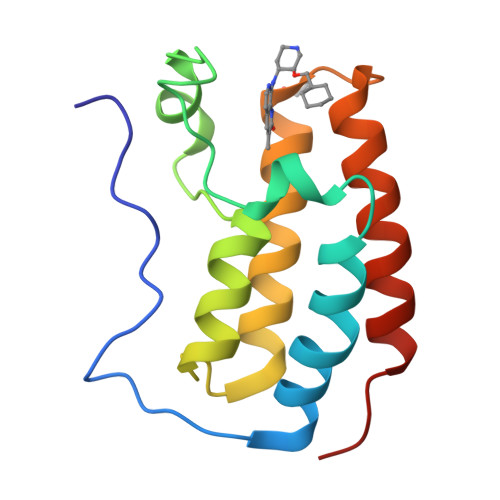
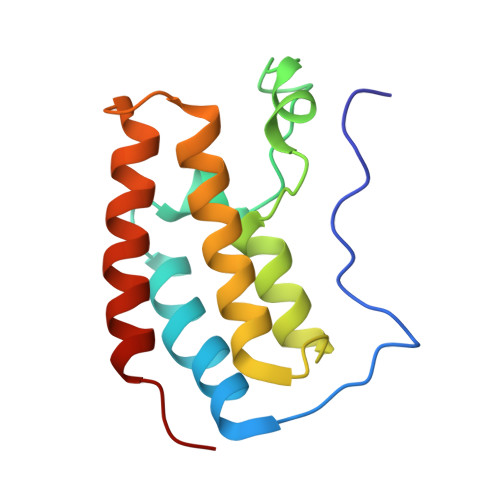
Structure-Based Optimization of Naphthyridones Into Potent Atad2 Bromodomain Inhibitors.
Bamborough, P., Chung, C., Furze, R.C., Grandi, P., Michon, A., Sheppard, R.J., Barnett, H., Diallo, H., Dixon, D.P., Douault, C., Jones, E.J., Karamshi, B., Mitchell, D.J., Prinjha, R.K., Rau, C., Watson, R.J., Werner, T., Demont, E.H.(2015) J Med Chem 58: 6151
- PubMed: 26230603
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00773
- Primary Citation of Related Structures:
5A81, 5A82, 5A83, 5A85 - PubMed Abstract:
ATAD2 is a bromodomain-containing protein whose overexpression is linked to poor outcomes in a number of different cancer types. To date, no potent and selective inhibitors of the bromodomain have been reported. This article describes the structure-based optimization of a series of naphthyridones from micromolar leads with no selectivity over the BET bromodomains to inhibitors with sub-100 nM ATAD2 potency and 100-fold BET selectivity.
Organizational Affiliation:
∥Cellzome GmbH, Molecular Discovery Research, GlaxoSmithKline, Meyerhofstrasse 1, 69117 Heidelberg, Germany.