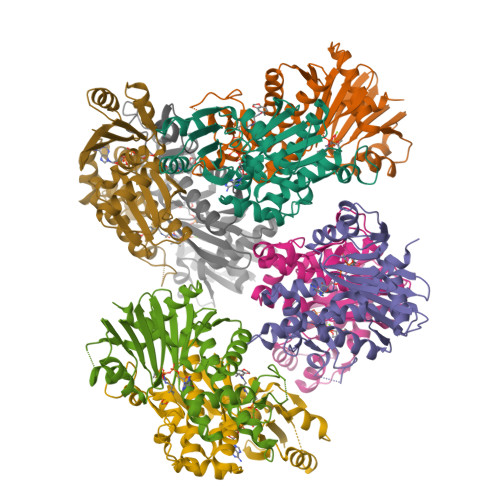
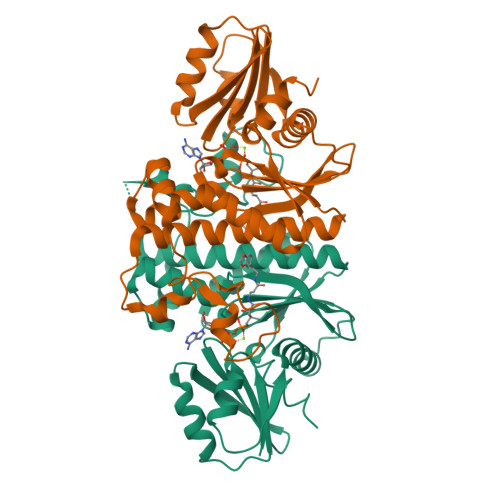
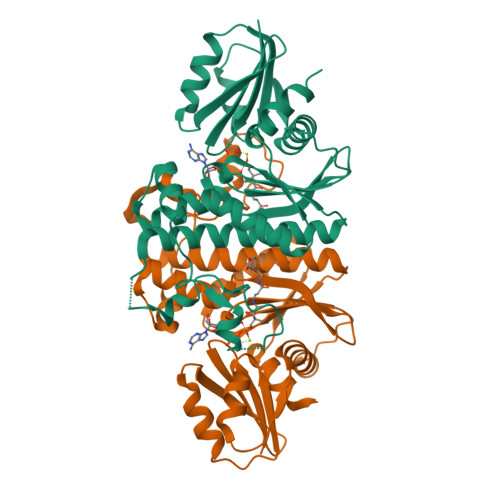
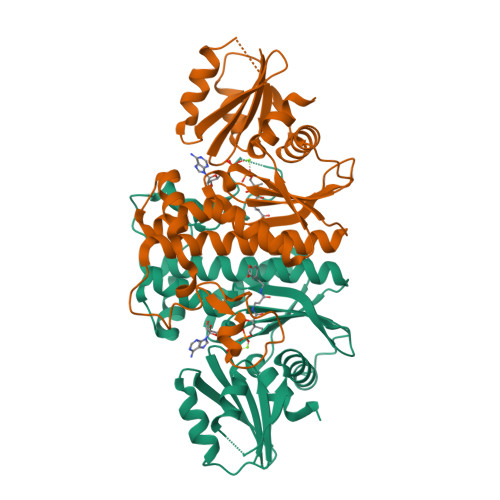
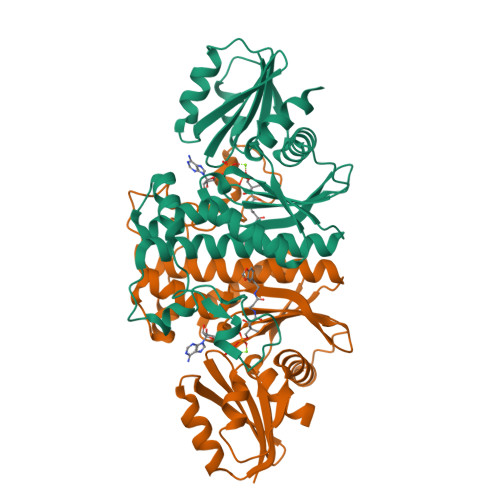
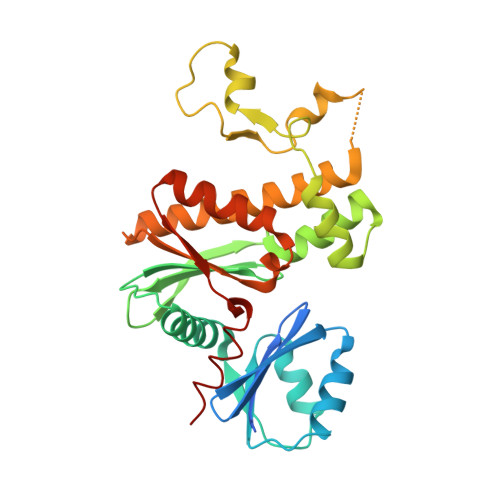
Structural characterization of a new N-substituted pantothenamide bound to pantothenate kinases from Klebsiella pneumoniae and Staphylococcus aureus.
Hughes, S.J., Antoshchenko, T., Kim, K.P., Smil, D., Park, H.W.(2014) Proteins 82: 1542-1548
- PubMed: 24470271
- DOI: https://doi.org/10.1002/prot.24524
- Primary Citation of Related Structures:
4NB4, 4NE2 - PubMed Abstract:
Pantothenate kinase (PanK) is the rate-limiting enzyme in Coenzyme A biosynthesis, catalyzing the ATP-dependent phosphorylation of pantothenate. We solved the co-crystal structures of PanKs from Staphylococcus aureus (SaPanK) and Klebsiella pneumonia (KpPanK) with N-[2-(1,3-benzodioxol-5-yl)ethyl] pantothenamide (N354-Pan). Two different N354-Pan conformers interact with polar/nonpolar mixed residues in SaPanK and aromatic residues in KpPanK. Additionally, phosphorylated N354-Pan is found at the closed active site of SaPanK but not at the open active site of KpPanK, suggesting an exchange of the phosphorylated product with a new N354-Pan only in KpPanK. Together, pantothenamides conformational flexibility and binding pocket are two key considerations for selective compound design.
Organizational Affiliation:
Department of Pharmacology, University of Toronto, Toronto, Ontario, M5G 1L7.