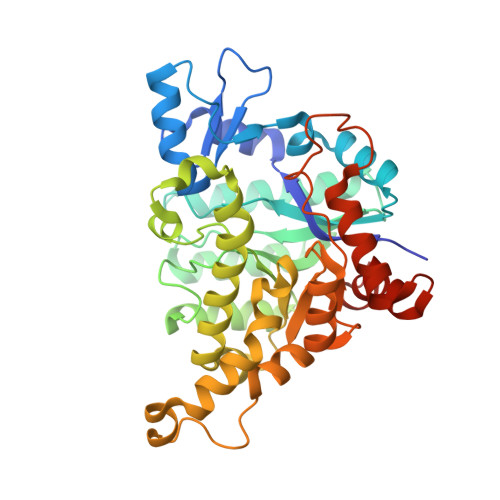
Crystal structures of isoorotate decarboxylases reveal a novel catalytic mechanism of 5-carboxyl-uracil decarboxylation and shed light on the search for DNA decarboxylase.
Xu, S., Li, W., Zhu, J., Wang, R., Li, Z., Xu, G.L., Ding, J.(2013) Cell Res 23: 1296-1309
- PubMed: 23917530
- DOI: https://doi.org/10.1038/cr.2013.107
- Primary Citation of Related Structures:
4HJW, 4HK5, 4HK6, 4HK7, 4LAK, 4LAL, 4LAM, 4LAN, 4LAO - PubMed Abstract:
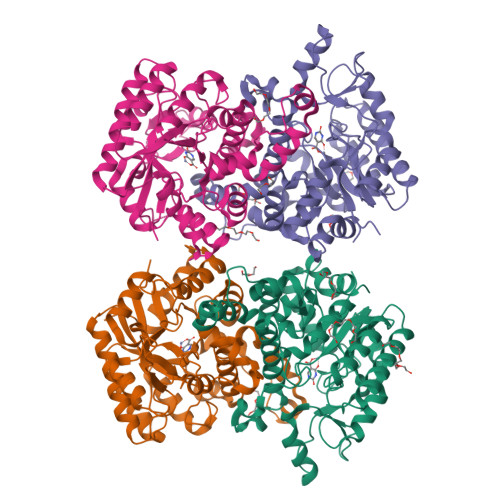
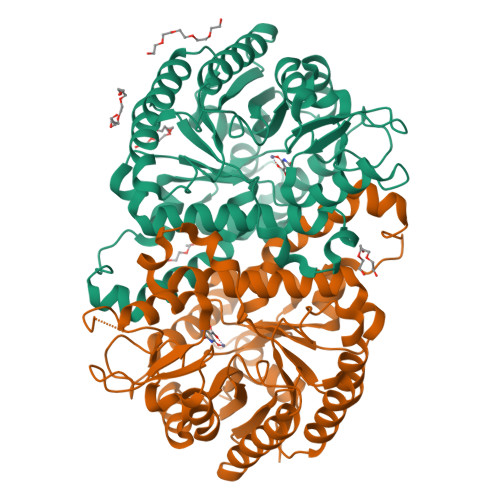
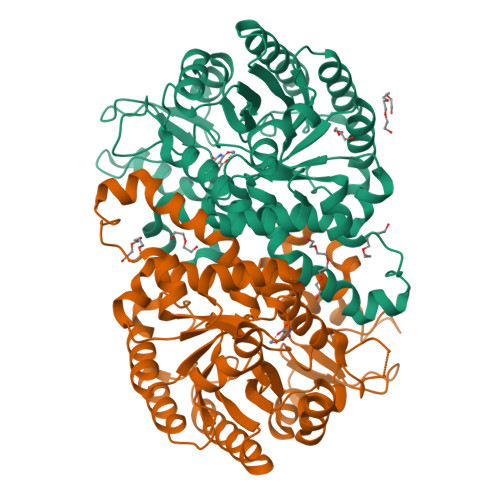
DNA methylation and demethylation regulate many crucial biological processes in mammals and are linked to many diseases. Active DNA demethylation is believed to be catalyzed by TET proteins and a putative DNA decarboxylase that may share some similarities in sequence, structure and catalytic mechanism with isoorotate decarboxylase (IDCase) that catalyzes decarboxylation of 5caU to U in fungi. We report here the structures of wild-type and mutant IDCases from Cordyceps militaris and Metarhizium anisopliae in apo form or in complexes with 5caU, U, and an inhibitor 5-nitro-uracil. IDCases adopt a typical (β/α)8 barrel fold of the amidohydrolase superfamily and function as dimers. A Zn(2+) is bound at the active site and coordinated by four strictly conserved residues, one Asp and three His. The substrate is recognized by several strictly conserved residues. The functional roles of the key residues at the active site are validated by mutagenesis and biochemical studies. Based on the structural and biochemical data, we present for the first time a novel catalytic mechanism of decarboxylation for IDCases, which might also apply to other members of the amidohydrolase superfamily. In addition, our biochemical data show that IDCases can catalyze decarboxylation of 5caC to C albeit with weak activity, which is the first in vitro evidence for direct decarboxylation of 5caC to C by an enzyme. These findings are valuable in the identification of potential DNA decarboxylase in mammals.
Organizational Affiliation:
1] State Key Laboratory of Molecular Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China [2] Graduate School of Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.