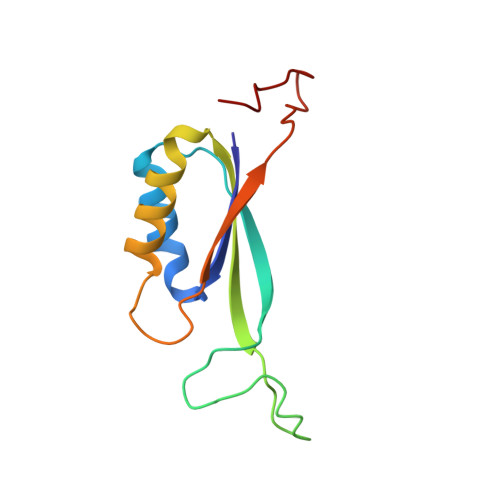
X-ray structure of the signal transduction protein from Escherichia coli at 1.9 A.
Carr, P.D., Cheah, E., Suffolk, P.M., Vasudevan, S.G., Dixon, N.E., Ollis, D.L.(1996) Acta Crystallogr D Biol Crystallogr 52: 93-104
- PubMed: 15299730
- DOI: https://doi.org/10.1107/S0907444995007293
- Primary Citation of Related Structures:
2PII - PubMed Abstract:
The structure of the bacterial signal transduction protein P(II) has been refined to an R factor of 13.2% using 3sigma data between 10 and 1.9 A. The crystals exhibited twinning by merohedry and X-ray intensities were corrected using the method of Fisher & Sweet [Fisher & Sweet (1980). Acta Cryst. A36, 755-760] prior to refinement. Our earlier 2.7 A structure [Cheah, Carr, Suffolk, Vasudevan, Dixon & Ollis (1994). Structure, 2, 981-990] served as a starting model. P(II) is a trimeric molecule, each subunit has a mass of 12.4 kDa and contains 112 amino-acid residues. The refined model includes all 1065 protein atoms per subunit plus 312 water molecules. The high-resolution refinement confirms the correctness of our 2.7 A model, although it leads to a redefinition of the extent of various secondary-structural elements. The monomeric structure of P(II) exhibits an interlocking double betaalphabeta fold. This is a stable fold found in a number of proteins with diverse functions. The association of the protein into a trimer leads to a new structure which we describe in detail. The effects of crystal packing forces are discussed and potential interaction sites with other proteins and effector molecules are identified.
Organizational Affiliation:
Centre for Molecular Structure and Function, Research School of Chemistry, Australian National University, Canberra, ACT.