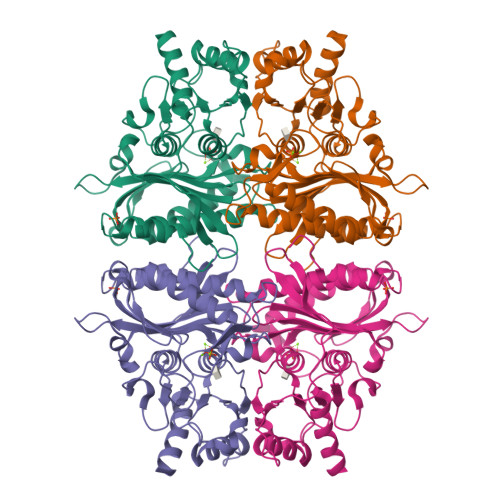
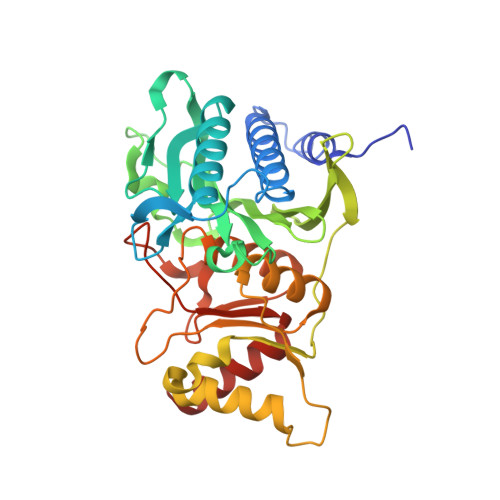
R-state AMP complex reveals initial steps of the quaternary transition of fructose-1,6-bisphosphatase.
Iancu, C.V., Mukund, S., Fromm, H.J., Honzatko, R.B.(2005) J Biol Chem 280: 19737-19745
- PubMed: 15767255
- DOI: https://doi.org/10.1074/jbc.M501011200
- Primary Citation of Related Structures:
1YXI, 1YYZ, 1YZ0 - PubMed Abstract:
AMP transforms fructose-1,6-bisphosphatase from its active R-state to its inactive T-state; however, the mechanism of that transformation is poorly understood. The mutation of Ala(54) to leucine destabilizes the T-state of fructose-1,6-bisphosphatase. The mutant enzyme retains wild-type levels of activity, but the concentration of AMP that causes 50% inhibition increases 50-fold. In the absence of AMP, the Leu(54) enzyme adopts an R-state conformation nearly identical to that of the wild-type enzyme. The mutant enzyme, however, grows in two crystal forms in the presence of saturating AMP. In one form, the AMP-bound tetramer is in a T-like conformation, whereas in the other form, the AMP-bound tetramer is in a R-like conformation. The latter reveals conformational changes in two helices due to the binding of AMP. Helix H1 moves toward the center of the tetramer and displaces Ile(10) from a hydrophobic pocket. The displacement of Ile(10) exposes a hydrophobic surface critical to interactions that stabilize the T-state. Helix H2 moves away from the center of the tetramer, breaking hydrogen bonds with a buried loop (residues 187-195) in an adjacent subunit. The same hydrogen bonds reform but only after the quaternary transition to the T-state. Proposed here is a model that accounts for the quaternary transition and cooperativity in the inhibition of catalysis by AMP.
Organizational Affiliation:
Department of Biochemistry, Biophysics, and Molecular Biology, Iowa State University, Ames, 50011, USA.