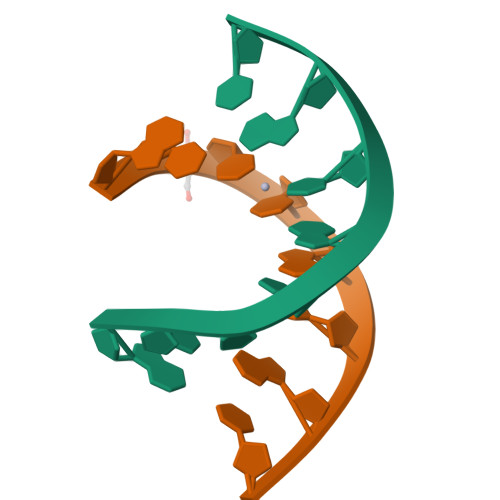
Flexibility and stabilization of HgII-mediated C:T and T:T base pairs in DNA duplex
Liu, H.H., Cai, C., Haruehanroengra, P., Yao, Q.Q., Chen, Y.Q., Yang, C., Luo, Q., Wu, B.X., Li, J.X., Ma, J.B., Sheng, J., Gan, J.H.(2017) Nucleic Acids Res 45: 2910-2918
- PubMed: 27998930
- DOI: https://doi.org/10.1093/nar/gkw1296
- Primary Citation of Related Structures:
5GSK, 5WSP, 5WSQ, 5WSR, 5WSS - PubMed Abstract:
Owing to their great potentials in genetic code extension and the development of nucleic acid-based functional nanodevices, DNA duplexes containing HgII-mediated base pairs have been extensively studied during the past 60 years. However, structural basis underlying these base pairs remains poorly understood. Herein, we present five high-resolution crystal structures including one first-time reported C-HgII-T containing duplex, three T-HgII-T containing duplexes and one native duplex containing T-T pair without HgII. Our structures suggest that both C-T and T-T pairs are flexible in interacting with the HgII ion with various binding modes including N3-HgII-N3, N4-HgII-N3, O2-HgII-N3 and N3-HgII-O4. Our studies also reveal that the overall conformations of the C-HgII-T and T-HgII-T pairs are affected by their neighboring residues via the interactions with the solvent molecules or other metal ions, such as SrII. These results provide detailed insights into the interactions between HgII and nucleobases and the structural basis for the rational design of C-HgII-T or T-HgII-T containing DNA nanodevices in the future.
Organizational Affiliation:
State Key Laboratory of Genetic Engineering, Collaborative Innovation Center of Genetics and Development, Department of Physiology and Biophysics, School of Life Sciences, Fudan University, Shanghai 200433, China.