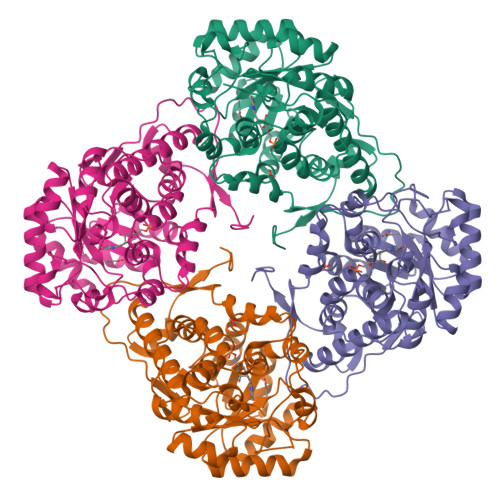
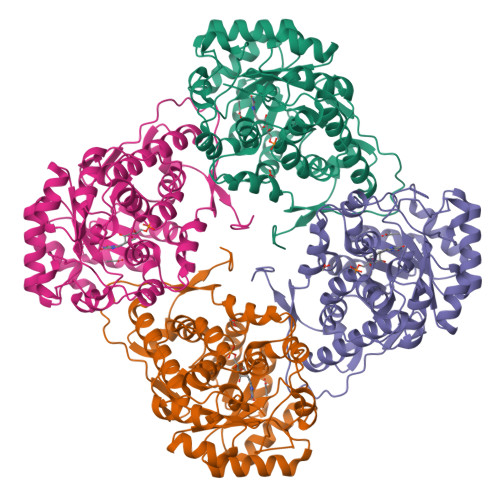
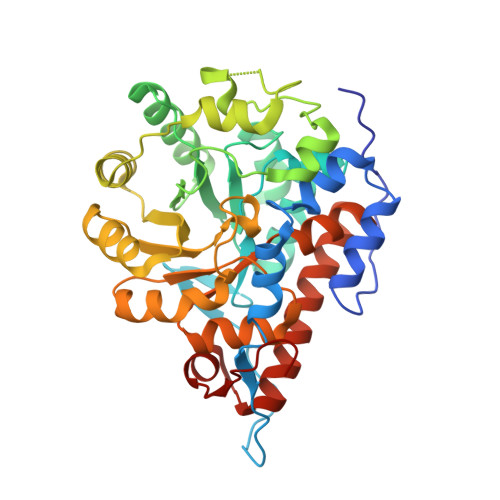
The Ala95-to-Gly substitution in Aerococcus viridans l-lactate oxidase revisited - structural consequences at the catalytic site and effect on reactivity with O2 and other electron acceptors.
Stoisser, T., Rainer, D., Leitgeb, S., Wilson, D.K., Nidetzky, B.(2015) FEBS J 282: 562-578
- PubMed: 25423902
- DOI: https://doi.org/10.1111/febs.13162
- Primary Citation of Related Structures:
4RJE - PubMed Abstract:
Aerococcus viridansl-lactate oxidase (avLOX) is a biotechnologically important flavoenzyme that catalyzes the conversion of L-lactate and O₂ into pyruvate and H₂O₂. The enzymatic reaction underlies different biosensor applications of avLOX for blood L-lactate determination. The ability of avLOX to replace O₂ with other electron acceptors such as 2,6-dichlorophenol-indophenol (DCIP) allows the possiblity of analytical and practical applications. The A95G variant of avLOX was previously shown to exhibit lowered reactivity with O₂ compared to wild-type enzyme and therefore was employed in a detailed investigation with respect to the specificity for different electron acceptor substrates. From stopped-flow experiments performed at 20 °C (pH 6.5), we determined that the A95G variant (fully reduced by L-lactate) was approximately three-fold more reactive towards DCIP (1.0 ± 0.1 × 10(6) M(-1) ·s(-1) ) than O₂, whereas avLOX wild-type under the same conditions was 14-fold more reactive towards O₂(1.8 ± 0.1 × 10(6) m(-1) ·s(-1)) than DCIP. Substituted 1,4-benzoquinones were up to five-fold better electron acceptors for reaction with L-lactate-reduced A95G variant than wild-type. A 1.65-Å crystal structure of oxidized A95G variant bound with pyruvate was determined and revealed that the steric volume created by removal of the methyl side chain of Ala95 and a slight additional shift in the main chain at position Gly95 together enable the accomodation of a new active-site water molecule within hydrogen-bond distance to the N5 of the FMN cofactor. The increased steric volume available in the active site allows the A95G variant to exhibit a similar trend with the related glycolate oxidase in electron acceptor substrate specificities, despite the latter containing an alanine at the analogous position.
Organizational Affiliation:
Research Center Pharmaceutical Engineering, Graz, Austria; Institute of Biotechnology and Biochemical Engineering, Graz University of Technology, NAWI Graz, Graz, Austria.