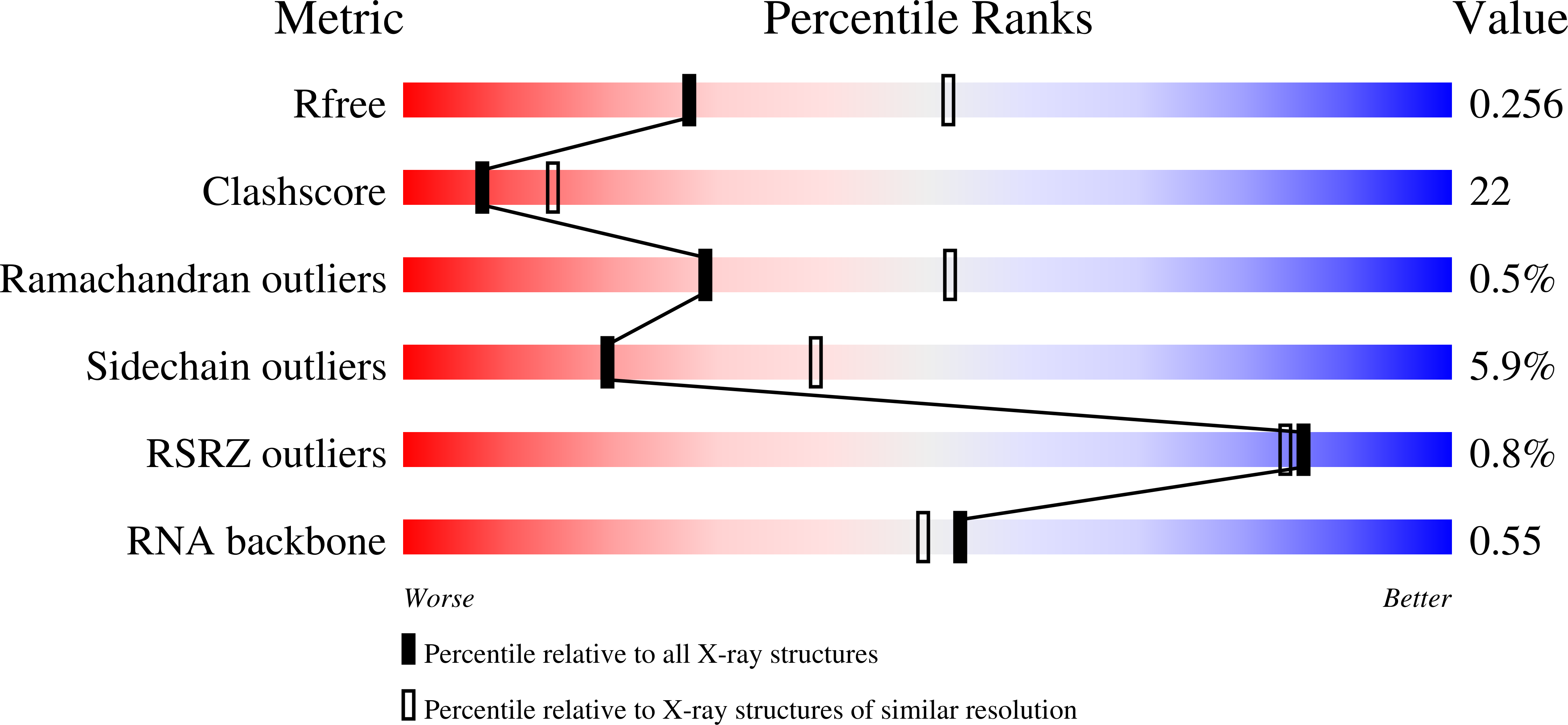
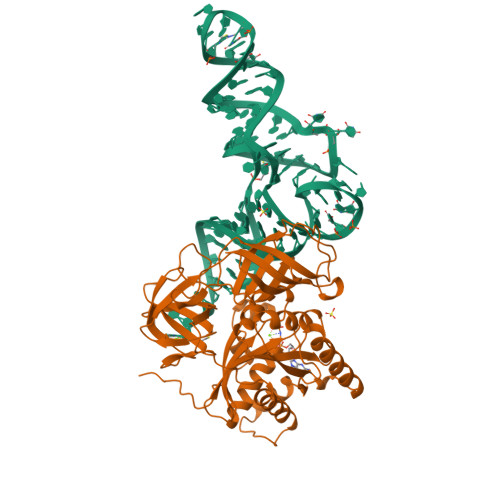
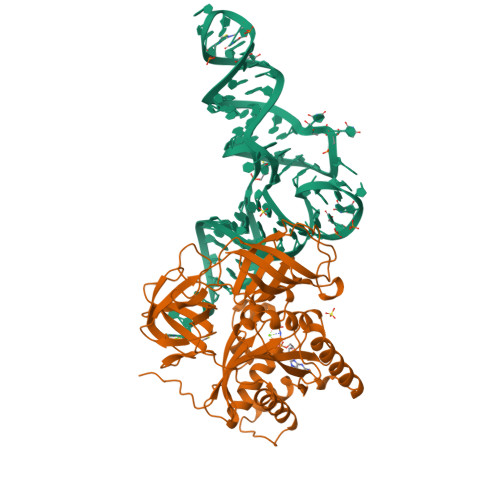
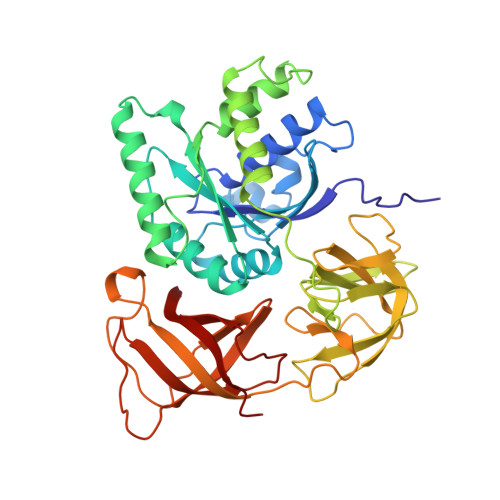
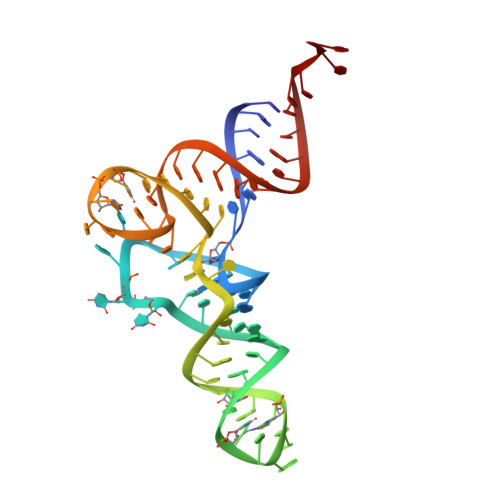
The crystal structure of Cys-tRNACys-EF-Tu-GDPNP reveals general and specific features in the ternary complex and in tRNA.
Nissen, P., Thirup, S., Kjeldgaard, M., Nyborg, J.(1999) Structure 7: 143-156
- PubMed: 10368282
- DOI: https://doi.org/10.1016/s0969-2126(99)80021-5
- Primary Citation of Related Structures:
1B23 - PubMed Abstract:
. The translation elongation factor EF-Tu in its GTP-bound state forms a ternary complex with any aminoacylated tRNA (aa-tRNA), except initiator tRNA and selenocysteinyl-tRNA. This complex delivers aa-tRNA to the ribosomal A site during the elongation cycle of translation. The crystal structure of the yeast Phe-tRNAPhe ternary complex with Thermus aquaticus EF-Tu-GDPNP (Phe-TC) has previously been determined as one representative of this general yet highly discriminating complex formation. The ternary complex of Escherichia coli Cys-tRNACys and T. aquaticus EF-Tu-GDPNP (Cys-TC) has been solved and refined at 2.6 degrees resolution. Conserved and variable features of the aa-tRNA recognition and binding by EF-Tu-GTP have been revealed by comparison with the Phe-TC structure. New tertiary interactions are observed in the tRNACys structure. A 'kissing complex' is observed in the very close crystal packing arrangement. The recognition of Cys-tRNACys by EF-Tu-GDPNP is restricted to the aa-tRNA motif previously identified in Phe-TC and consists of the aminoacylated 3' end, the phosphorylated 5' end and one side of the acceptor stem and T stem. The aminoacyl bond is recognized somewhat differently, yet by the same primary motif in EF-Tu, which suggests that EF-Tu adapts to subtle variations in this moiety among all aa-tRNAs. New tertiary interactions revealed by the Cys-tRNACys structure, such as a protonated C16:C59 pyrimidine pair, a G15:G48 'Levitt pair' and an s4U8:A14:A46 base triple add to the generic understanding of tRNA structure from sequence. The structure of the 'kissing complex' shows a quasicontinuous helix with a distinct shape determined by the number of base pairs.
Organizational Affiliation:
Institute of Molecular and Structural Biology, Aarhus University, Gustav Wieds Vej 10, C DK 8000 Aarhus C, Denmark.