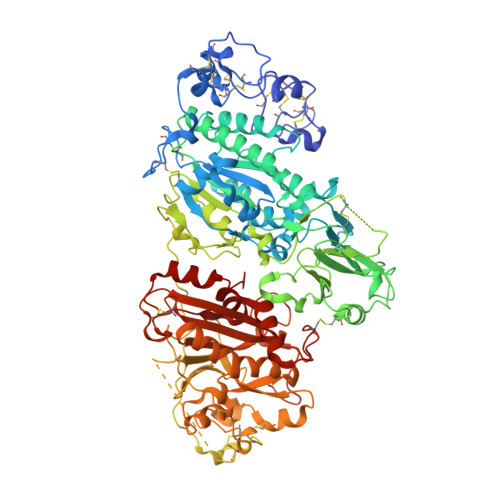
Structural Basis for Inhibition of Human Autotaxin by Four Potent Compounds with Distinct Modes of Binding.
Stein, A.J., Bain, G., Prodanovich, P., Santini, A.M., Darlington, J., Stelzer, N.M., Sidhu, R.S., Schaub, J., Goulet, L., Lonergan, D., Calderon, I., Evans, J.F., Hutchinson, J.H.(2015) Mol Pharmacol 88: 982-992
- PubMed: 26371182
- DOI: https://doi.org/10.1124/mol.115.100404
- Primary Citation of Related Structures:
4ZG6, 4ZG7, 4ZG9, 4ZGA - PubMed Abstract:
Autotaxin (ATX) is a secreted enzyme that hydrolyzes lysophosphatidylcholine to lysophosphatidic acid (LPA). LPA is a bioactive phospholipid that regulates diverse biological processes, including cell proliferation, migration, and survival/apoptosis, through the activation of a family of G protein-coupled receptors. The ATX-LPA pathway has been implicated in many pathologic conditions, including cancer, fibrosis, inflammation, cholestatic pruritus, and pain. Therefore, ATX inhibitors represent an attractive strategy for the development of therapeutics to treat a variety of diseases. Mouse and rat ATX have been crystallized previously with LPA or small-molecule inhibitors bound. Here, we present the crystal structures of human ATX in complex with four previously unpublished, structurally distinct ATX inhibitors. We demonstrate that the mechanism of inhibition of each compound reflects its unique interactions with human ATX. Our studies may provide a basis for the rational design of novel ATX inhibitors.
Organizational Affiliation:
Cayman Chemical Company, Ann Arbor, Michigan (A.J.S., N.M.P.S., R.S.S., J.S.); and PharmAkea, San Diego, California (G.B., P.P., A.M.S., J.D., L.G., D.L., I.C., J.F.E., J.H.H.).