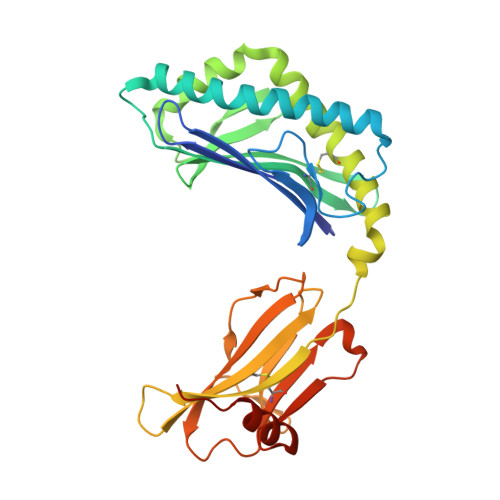
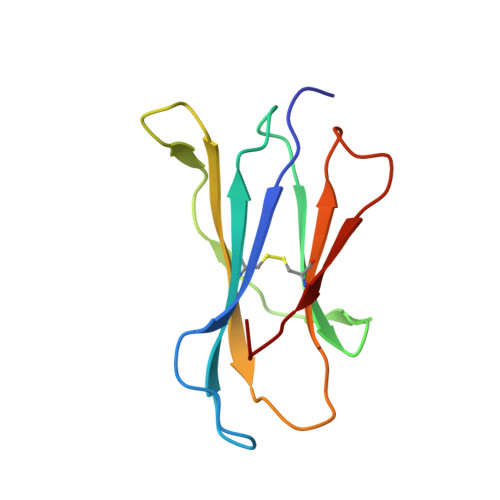
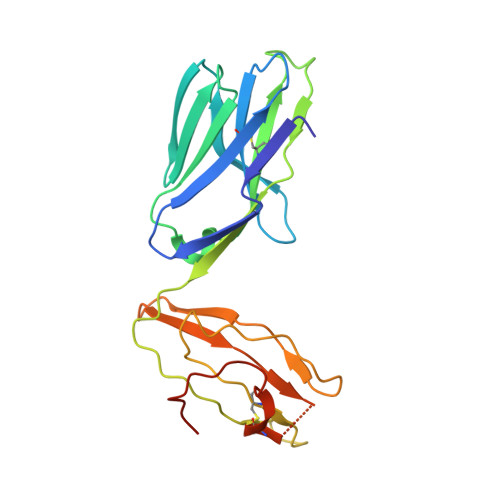
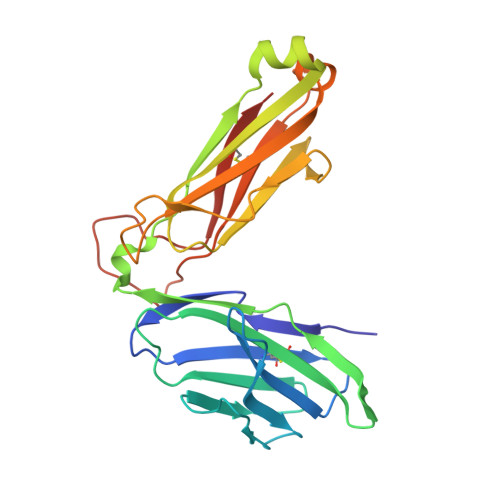
A molecular basis for NKT cell recognition of CD1d-self-antigen
Mallevaey, T., Clarke, A.J., Scott-Browne, J.P., Young, M.H., Roisman, L.C., Pellicci, D.G., Patel, O., Vivian, J.P., Matsuda, J.L., McCluskey, J., Godfrey, D.I., Marrack, P., Rossjohn, J., Gapin, L.(2011) Immunity 34: 315-326
- PubMed: 21376640
- DOI: https://doi.org/10.1016/j.immuni.2011.01.013
- Primary Citation of Related Structures:
3AU1, 3QI9 - PubMed Abstract:
The antigen receptor for natural killer T cells (NKT TCR) binds CD1d-restricted microbial and self-lipid antigens, although the molecular basis of self-CD1d recognition is unclear. Here, we have characterized NKT TCR recognition of CD1d molecules loaded with natural self-antigens (Ags) and report the 2.3 Å resolution structure of an autoreactive NKT TCR-phosphatidylinositol-CD1d complex. NKT TCR recognition of self- and foreign antigens was underpinned by a similar mode of germline-encoded recognition of CD1d. However, NKT TCR autoreactivity is mediated by unique sequences within the non-germline-encoded CDR3β loop encoding for a hydrophobic motif that promotes self-association with CD1d. Accordingly, NKT cell autoreactivity may arise from the inherent affinity of the interaction between CD1d and the NKT TCR, resulting in the recognition of a broad range of CD1d-restricted self-antigens. This demonstrates that multiple self-antigens can be recognized in a similar manner by autoreactive NKT TCRs.
Organizational Affiliation:
Department of Immunology, University of Colorado School of Medicine and National Jewish Health, Denver, CO 80206, USA.